|
In this podcast, I chat with Matt and Hanna at 80/20 endurance
I’ve learned a lot over the years from Matt’s books, and it was a real pleasure to spend some time talking about HRV and training Thank you & enjoy I have recently joined the editorial board of IEEE Pervasive Computing Magazine.
Here you can find our first article, written with Lucy Dunne, and covering future trends in wearable technology Enjoy the read The Use Case for Heart Rate Variability with Marco Altini and Corrine Malcolm | Koopcast Episode 1069/12/2021
Last week I had the opportunity to chat with Jason Koop and Corrine Malcom about heart rate variability (HRV), "readiness" scores, resting physiology, and wearables, in the context of training
If you are interested in understanding the nuances of these aspects, including their strengths and limitations, this episode is a good starting point Don't fall for cults or for those who trivialize everything that is human physiology. Instead, strive to understand these aspects, and you'll be a better athlete or coach Thanks again Jason! Excited to share our latest paper on heart rate and heart rate variability (HRV):
This one has been in the making for a long time, and brings it all together: developing an accurate and affordable method to measure resting physiology, analyzing data at a scale that goes far beyond what we can do in the lab, and as a result, new, valuable insights Give it a read, here Professor Maria Carrasco-Poyatos and co-authors at the University of Almería just published the latest paper on HRV-guided training, titled "Heart rate variability-guided training in professional runners: effects on performance and vagal modulation"
Thank you Maria and co-authors for using HRV4Training and for involving me in this work. You can find the paper, here, including the main highlights:
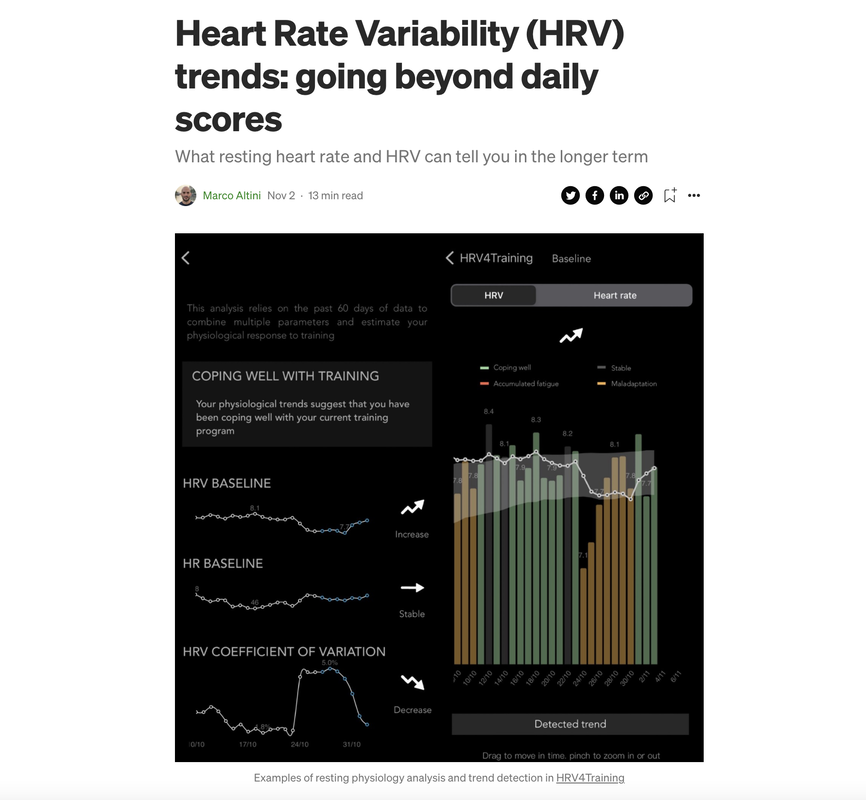
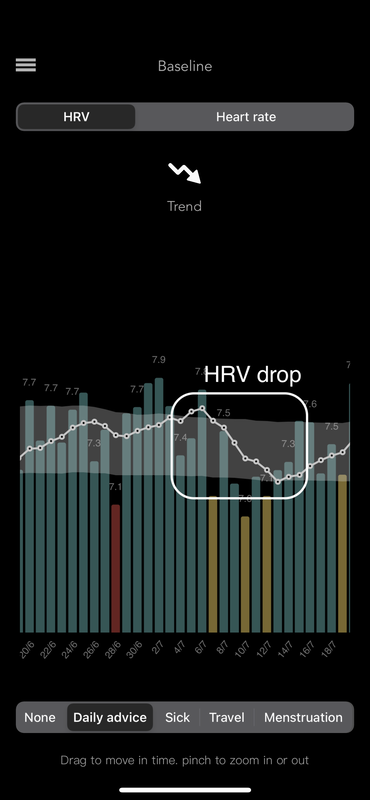
Heart rate variability (HRV) trends over long periods of time (e.g. from weeks to months) are one of the most interesting and complex aspects to analyze when it comes to resting physiology
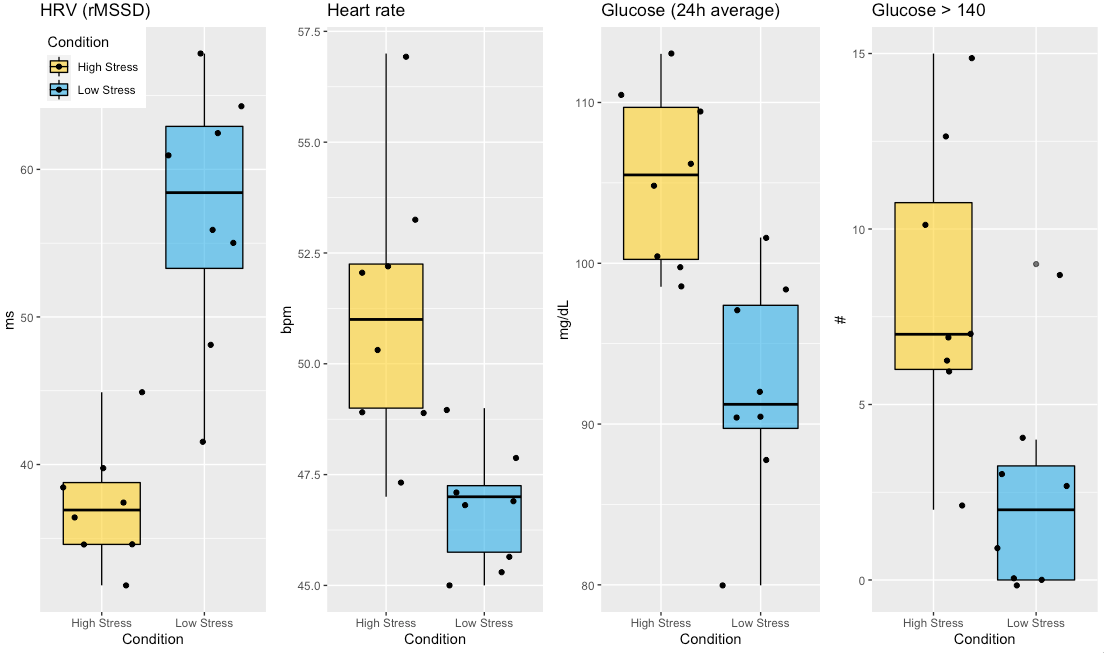
While day-to-day (or acute) changes reflect well stressors such as training intensity, the menstrual cycle, sickness, alcohol intake, or travel in the day(s) before the measurement, in the long term things are quite different In this post, I will cover our approach to trends analysis in HRV4Training, and cover some of the features in the app that should help you make sense of the data in the longer term Learn more, here How does the body respond to stress? Below I look at heart rate variability (HRV), heart rate, and glucose in response to two very different weeks (N = 1). High vs low stress:
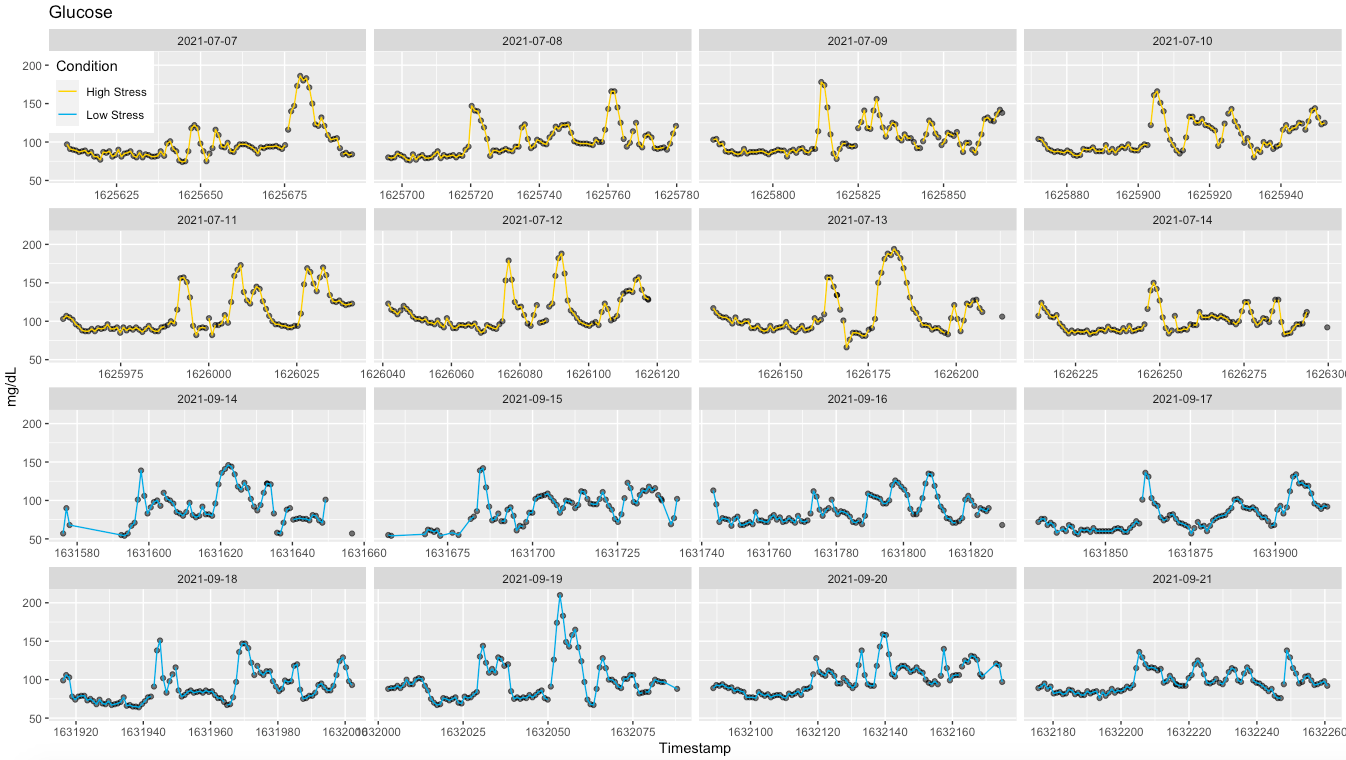
Some context first Last summer in July I had a strong negative stress response (cumulative stressors), resulting in arrhythmia and concerns for my health I've talked about this before, but here I want to focus on what happened to glucose during that week. Coincidentally, I was wearing a continuous glucose monitor (CGM) since the previous week and noticed that after meals, my glucose was spiking really high, near 200 mg/dL, consistently. Very interesting to see poor regulation at work so clearly. As usual, I was also monitoring my resting physiology (HRV and HR) using HRV4Training (morning measurements), and saw quite a dip in HRV, as well as a minor change in heart rate This is the type of stress response I often discuss (see for example my guide here) Physiologically, we know that high stress is associated acutely and chronically with elevated glucose in the bloodstream and reduced parasympathetic activity Pretty neat to see it with simple measurements and currently available technology.
"If you've ever wondered how to tap into the secrets of how that pump in your chest can help you to train faster, harder and longer Marco is just the man to listen to."
“We've been using Marco's app for a while to understand how to better regulate training and recovery and in this episode we do a deep dive on the how and why” I’ve really enjoyed talking to ultrarunners Jay and Tris about HRV, thanks for having me! Episode here.
In this interview, we cover:
Thank you Kieran for having me on your channel Excited to announce that I am joining the editorial board of IEEE Pervasive Computing Magazine
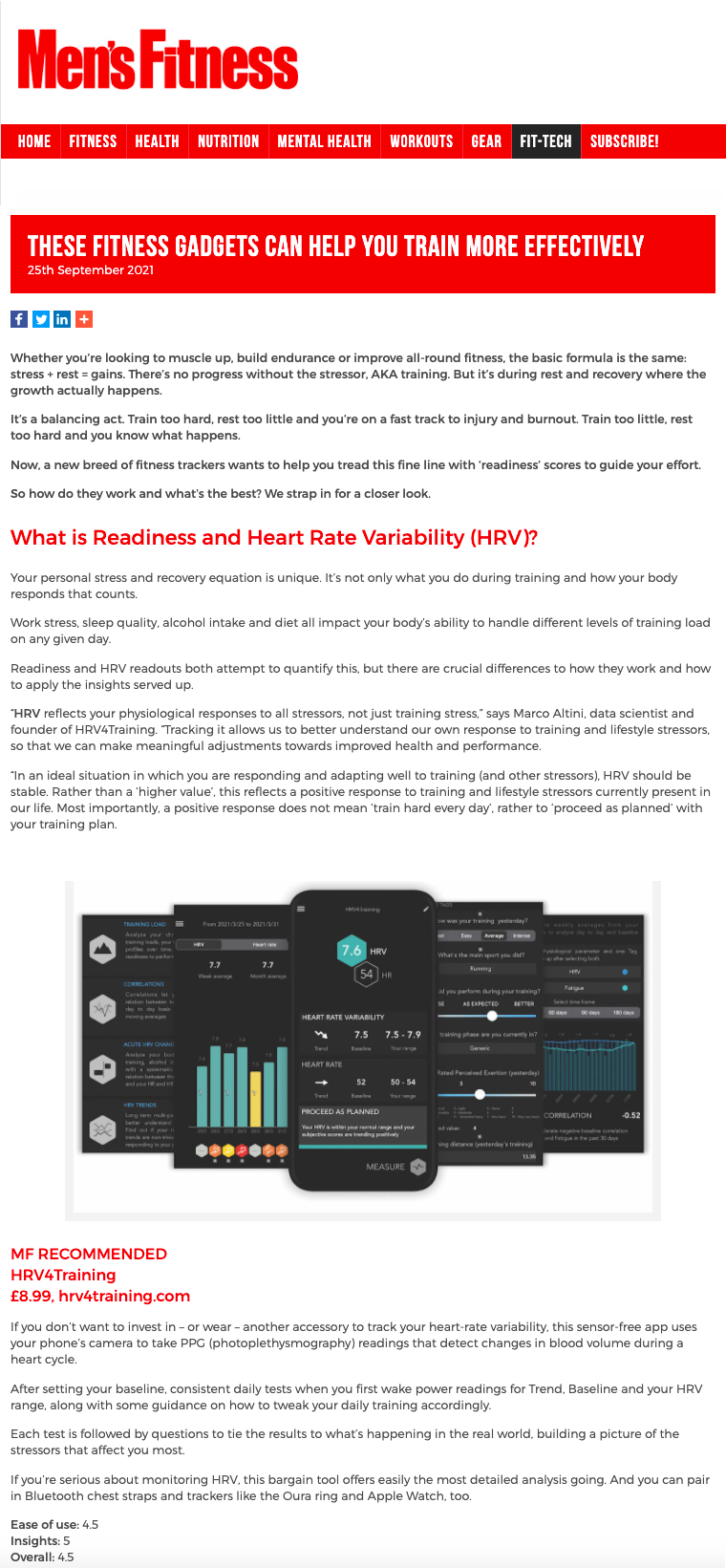
I will take a role as editor for the Wearables Department together with Lucy Dunne Our first editorial should be out soon sometime soon “HRV reflects your physiological responses to all stressors, not just training stress,” says Marco Altini
“Tracking HRV allows us to better understand our own response to training and lifestyle stressors, so that we can make meaningful adjustments towards improved health and performance" Thank you Men's Fitness Mag for featuring HRV4Training and ŌURA. Find the article, here
Last week I had a nice chat with Dr. Greg Wells for his podcast. We talked wearables, validity, what’s measured (HRV), what’s estimated (sleep), how to use HRV data, and more
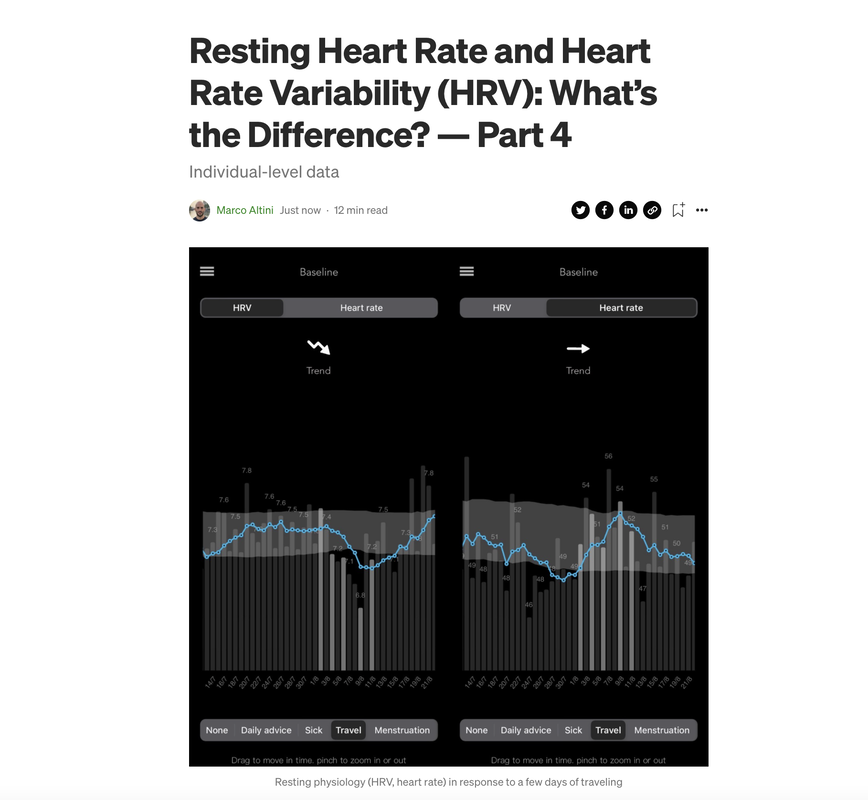
You can find the episode here. Thanks again Greg for having me In part 1 of this series, I covered the basic physiology of heart rhythm regulation. In part 2, I discussed the technology required for these measurements, why some sensors can be trusted, and why others can be used just for resting heart rate, and not for HRV. In part 3, we started looking at the data, with an analysis of population-level differences in resting heart rate and HRV.
In this blog, we finally get to the most interesting aspect: individual-level data. Needless to say, both resting heart rate and HRV become a lot more useful when tracked over time within individuals, and this is exactly what I’ll be showing here. I’ll also try to highlight some of the differences between these two parameters, so that you can better understand what the data means when tracked in response to strong acute stressors (e.g. training, sickness, alcohol intake, the menstrual cycle) and in the longer run (e.g. changes in fitness). You can find the blog, here. The goal of this post is to provide some clarity and general considerations on heart rate variability (HRV), readiness and wearables. I will try to clarify why comparing HRV and readiness scores is of little use and what you should be comparing (if anything) for a more meaningful assessment of how these devices work. Most importantly, we will see how you can benefit from the data for both HRV and readiness. What are we talking about?HRV is a measure of physiological stress. For today's wearables and apps, it typically represents parasympathetic activity due to how it is measured (at rest, while sleeping or first thing in the morning) and computed (relying on high frequency changes captured by rMSSD). This means that a lower HRV with respect to your historical data, is associated with higher stress. Readiness is a made up construct that most apps or wearables provide. The goal of readiness is to combine multiple parameters (one of them typically is HRV), to determine your level of recovery or ability to tackle the day (whatever that means in your case). Why does this matter?Due to the novelty of some of these metrics for consumers, issues in science communication, and whatnot, there is much confusion on either of them, to the point that often I see people comparing HRV from one wearable with readiness from another. While understandable (the tools are supposed to do the same thing, measure our recovery), this is like comparing apples with pears, it does not make much sense.
This is an important aspect to address because wearables and apps can be extremely helpful in better understanding physiological responses to the various stressors we face, but not all devices are equal, nor differences between the output of one or the other device necessarily mean that they cannot be trusted. Part 1 of my latest guide is all about physiology:
‣ Why do we care ‣ Bird’s-eye view ‣ Understanding autonomic control of the heart and more I hope you'll find it useful, enjoy the read HRV4Training just got a new look (both on Android and iPhone)
While we have made quite a few changes, some of the most important are in the homepage, which now displays:
Learn more about the only independently validated, camera-based HRV app, in our QuickStart guide: https://www.hrv4training.com/quickstart-guide.html which covers the basics of HRV, how to use the data, the various insights present in the app & more We hope you'll enjoy the new interface, and thank you for your support Our latest paper was just accepted for publication in Frontiers in Sports and Active Living: Elite Sports and Performance Enhancement.
In this paper, we show a case study of our real-time implementation of DFA alpha-1 in the HRV Logger, which you can find at this link. Learn more about the paper, here |
Marco ALtiniFounder of HRV4Training, Advisor @Oura , Guest Lecturer @VUamsterdam , Editor @ieeepervasive. PhD Data Science, 2x MSc: Sport Science, Computer Science Engineering. Runner Archives
May 2023
|








 RSS Feed
RSS Feed